Identification, also called supervised learning or classification, is no doubt one of the most important techniques in bioinformatics. The possibility to identify unknown organisms based upon various available experiment data sets is also a big step forward realized in BIONUMERICS, leading to more faithful consensus identifications. The same range of similarity and distance coefficients available for cluster analysis can be used for identification. In addition, state-of-the-art classifiers such as Naive Bayesian Classifiers and Support Vector Machines can be used.
Classifiers
Classifiers are the algorithms that perform identification or classification, i.e. the process of assigning biological samples to classes (categories) based on a training set of which the class membership is already known. The samples are analyzed based on experimental data, which can be quantitative, binary or categorical.
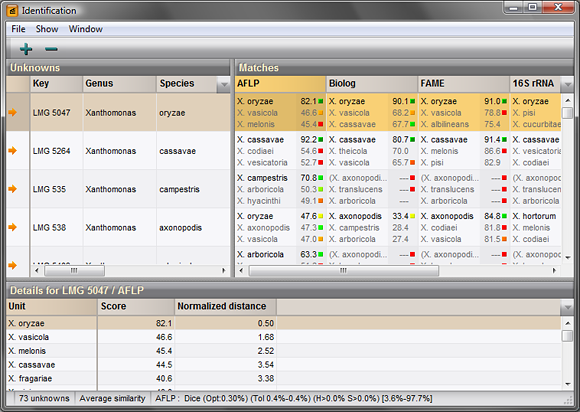
Database screening
Entire BIONUMERICS databases or selections thereof can be screened using fast matching algorithms to identify batches of entries based on fingerprint, character or sequence experiments. All available similarity or distance coefficients can be used for the fast screening. Matching entries are arranged by decreasing distance with the unknowns and can be selected for further comparison e.g. using cluster analysis.
Decision networks
Decision networks allow automated workflows to be built that make decisions, predict features, perform queries, fill in fields, create graphs and plots, etc. They can be used for resistance prediction, in breeding research, for complex reporting or for automated analysis of multilevel and polyphasic data analysis and data sorting. It includes the option to build bifurcating decision trees.
