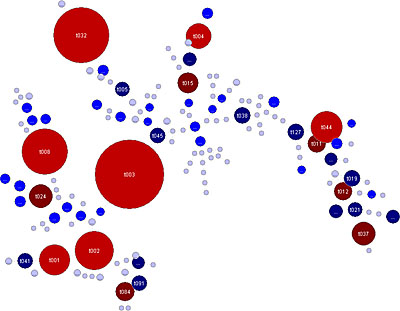
Variable-Number Tandem Repeats (VNTR's) are well known for their high mutation rate and are therefore widely used for subtyping. Some VNTR's, however, exhibit polymorphism in their individual repeat sequences. By sequencing the polymorphic repeat region, each new repeat variant determined can be assigned a unique repeat code. The repeat succession for a given strain in turn, determines that strain's VNTR type.
Polymorphic VNTR regions have been explored in various infectious bacteria for quick, portable, and inexpensive subtyping in hospital and outbreak settings. The best known example is undoubtedly spa typing, which is widely used for subtyping of Staphylococcus aureus. Because of the specificity of spa typing in terms of standardization, server synchronization and various other settings, a separate spa typing plugin is available in BIONUMERICS.
Below is an example of dru typing, a polymorphic VNTR in S. aureus.
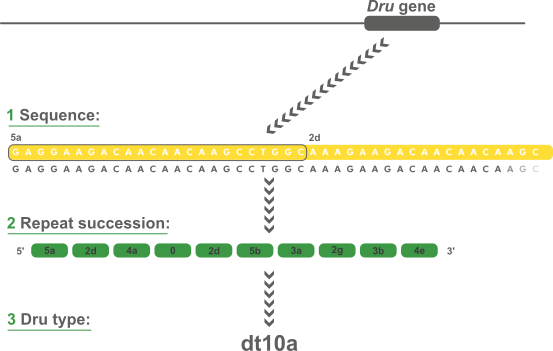
Polymorphic VNTR sequence typing in BIONUMERICS
This plugin offers the same functionality as the spa typing plugin. However, whereas the spa typing plugin is designed for routine analyses using a standardized workflow, the polymorphic VNTR typing plugin is rather a research tool that allows (and requires) various settings to be defined by the user. These include the name of the VNTR, the start and stop trimming positions, and the definitions for repeat and repeat succession types (file or URL based). Multiple polymorphic VNTR's can be defined, compared and analyzed in a combined way for the same organism.
BIONUMERICS provides a fully automated workflow, from import of raw sequencer trace files to assignment of repeat codes and repeat succession types. All problems or warnings during the workflow are reported and can be addressed with a single mouse click. BIONUMERICS can automatically synchronize repeat and repeat succession type signatures with a server, if defined. Or in a research phase, the user can add types manually.
Furthermore, BIONUMERICS offers a rich databasing and analysis platform, where multiple polymorphic VNTR sequence typing data sets can be compared, clustered and analyzed together or in conjunction with other subtyping data such as MLST, PFGE and any other phenotypic, genotypic, epidemiologic and geographic data.
The BIONUMERICS polymorphic VNTR typing plugin has been used for the development of several VNTR-based typing techniques, for example TR6 and TR10 typing in Clostridium difficile (see BMC Microbiology 2009, 9:6) and dru typing in Staphylococcus aureus.
Easy database setup
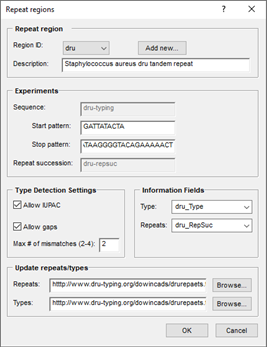 When you installing the plugin, BIONUMERICS allows you to define the VNTR's to be used, the trimming positions for each VNTR, the database fields, settings for assembly, and optionally, the files or server URL(s) for repeat and repeat succession types:
When you installing the plugin, BIONUMERICS allows you to define the VNTR's to be used, the trimming positions for each VNTR, the database fields, settings for assembly, and optionally, the files or server URL(s) for repeat and repeat succession types:
- Sequence and character experiments are created for the VNTR sequences and repeat successions, respecitively.
- Start and stop trimming positions are stored in the assembly template but can be changed by the user whenever appropriate.
- Base calling quality control settings (PHRED based) can be applied if required by an upload server.
- Information fields are created for repeat successions and VNTR types.
- Server URLs for repeats and types can be defined and changed anytime by the user if the server's URL changes.
- The software automatically downloads the repeat and type definitions at installation, and anytime at the user's request.
Full automatic processing workflow
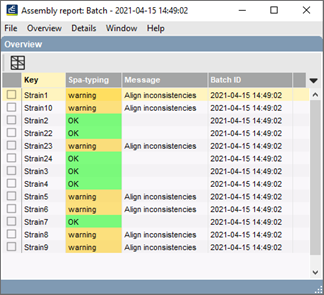
- Automatic import and assembly of batches of sequencer trace files from various sources (AB, Beckman, Amersham, FASTA); file names are parsed into strain and gene information using a parsing definition.
- Consensus sequences are automatically trimmed using start and stop signatures and placed in the right direction.
- When the batch assembly is finished, an overview report is shown, listing status of each strain/gene combination.
- Double-click on a problem contig to display the detailed information window.
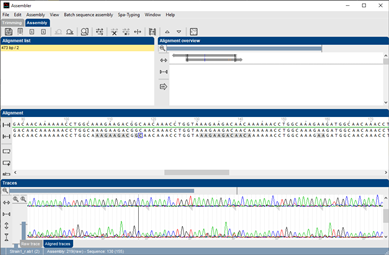
- Double-click on a particular problem to open the Assembler with the problem position selected.
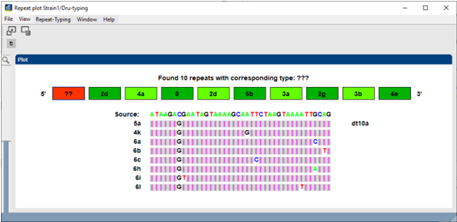
- For each unknown or problem repeat, show nearest existing repeats and the differences in bases - easy verification with chromatograms.
- Repeat successions are instantly identified as VNTR types using the synchronized database.
- For new (non-existing) repeat successions, the user can define types manually.
- Calculate population modelling networks in the finest and most comprehensive cluster analysis application available today, using the DSI alignment model and Minimum Spanning Trees or phylogenetic clustering methods.
- Calculate and display partitioning for clonal complexes and use BIONUMERICS' rich set of statistics tools.