The sequence read sets experiment type offers an integrated environment for importing, preprocessing and analyzing sets of reads from high throughput sequencers or public repositories.
An integrated NGS data analysis platform
- Fast import of sequence read sets from various next generation sequencing platforms, such as Roche 454, Illumina Solexa, IonTorrent, etc.
- Storage of large amounts of short sequences (including paired-end reads) and quality scores.
- Comprehensive data preprocessing and quality control settings for demultiplexing, splitting paired-end reads, primer removal, structural and quality trimming, chimera detection and cleaning up sequence read sets.
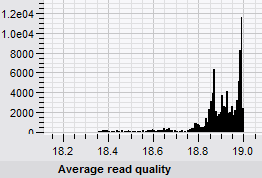
- Global statistics calculation of sequence reads: creation of read length histograms, revision of base distribution, and quality score distribution. Generation of reports in rich text, table and chart formats.
- Sequence read sets are database objects, meaning that they can be annotated using custom information fields and that user privileges determine who is allowed to access and/or modify the data.
- Create comparisons for Kmer based clustering of sequence read sets, using all available similarity coefficients and hierarchical clustering methods.
Supported in BIONUMERICS configurations:
BIONUMERICS-SEQ
BIONUMERICS-SUITE
See movie:
See tutorial:
