BIONUMERICS features a powerful tool for processing capillary sequencer fingerprints, i.e. electropherograms obtained from fragment analysis on multichannel capillary electrophoresis (CE) equipment.
The processing of a full capillary electrophoresis run can now be completed in seconds, rather than in minutes and consists of the following three steps:
- Import of electropherograms
- Peak detection
- Normalization
Electropherogram import
Intuitive import of raw electropherograms generated by Applied Biosystems (life technologies) genetic analyzers (3730xL, 3730, 3500xL, etc.) and Beckman Coulter CE equipment such as the GenomeLab™ GeXP Genetic Analysis System. Sample names can be parsed from the file names and data from different fluorescent channels are imported into the corresponding fingerprint type experiments.
Visualization
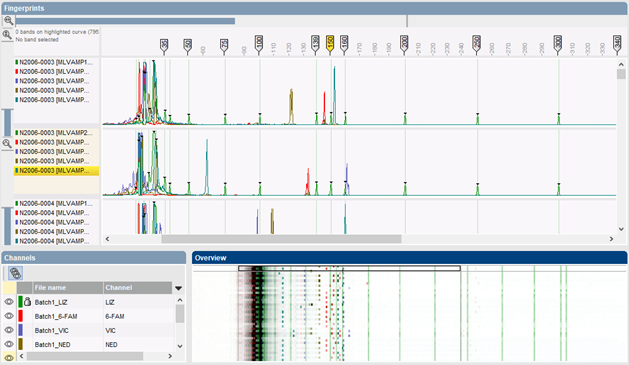
All electropherograms from the same CE run are uploaded in the Curve processing window and visualized as horizontal curves or pseudo-gel lanes. The fluorescent channels can be shown separate or in any combination. The window furthermore provides easy zooming and navigation tools.
Peak detection and filtering
The flexible peak detection algorithm employs user-defined thresholds and optional peak filters, specifically designed for CE profiles.
Stutter peaks and other artifacts
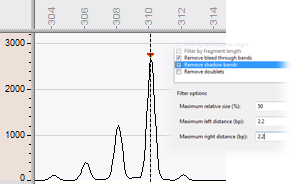 A specific stutter peak (or shadow peak) filter eliminates stutter bands, typically occurring in Variable Number of Tandem Repeats (VNTR) or Microsatellite analyses. Adjustable filter options allow to fine-tune the filter action. Spikes, curve noise and peak doublets are removed with their own dedicated filters.
A specific stutter peak (or shadow peak) filter eliminates stutter bands, typically occurring in Variable Number of Tandem Repeats (VNTR) or Microsatellite analyses. Adjustable filter options allow to fine-tune the filter action. Spikes, curve noise and peak doublets are removed with their own dedicated filters.
Dealing with bleed-through (pull-up peaks)
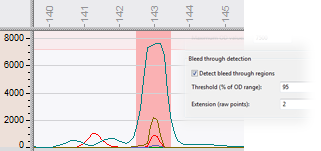 Bleed-through or pull-up is a typical problem in multichannel CE, which is caused by fluorescent dye cross-talk. A high signal in one fluorescent channel thereby gives rise to a pull-up peak in another channel. The software visualizes the bleed-through areas in a customizable background color and employs a filter to eliminate the pull-up peaks.
Bleed-through or pull-up is a typical problem in multichannel CE, which is caused by fluorescent dye cross-talk. A high signal in one fluorescent channel thereby gives rise to a pull-up peak in another channel. The software visualizes the bleed-through areas in a customizable background color and employs a filter to eliminate the pull-up peaks.
Easy normalization
Internal size markers are typically used in capillary electrophoresis, which allows for a very accurate normalization. A size marker can be simply picked from a list, for automatic generation of a reference system. The list currently contains 12 commercially available nucleic acid ladders, including the popular GeneScan™ 1200 LIZ® marker, featuring 68 DNA fragments.
An advanced normalization algorithm, that was specifically developed for CE profiles, reliably assigns marker peaks to reference positions. BIONUMERICS' well-known distortion bars feature is used to assess the quality of the thus-obtained normalization.
Compatible with existing data
Although the Curve processing window is completely new and a great time saver, by no means it represents a break with the past: data analyzed with the new tool are fully compatible with existing AFLP, MLVA, MLPA, SSR, etc. databases in BIONUMERICS. In fact, the user can even choose with processing tool to use: the new Curve processing window or the traditional Gel processing window. However, given its numerous advantages, we expect that most (if not all) CE users will embrace the new curve processing tool!
